We looked previously at data structures like vectors, and data types or modes, as in numeric, character string or logical. I’ll recap some of the previous post on vectors and then move on to talking about data structures like matrices (this post), and data frames (in brief here, in detail next post). This is because where we’re going we’ll need to know about things, and where we’re going is to the capacity to be able to create and manipulate data structures. (Other data structures, arrays and lists, will appear later.)
Data structures
The entities that R creates and manipulates are known as objects. These may be variables, arrays of numbers, character strings, functions, or more general structures built from such components.
During an analysis, an object is created, stored and used by name.
You can create and use objects – data structures – such as scalars, vectors, matrices, arrays, dataframes, and lists.
Vectors
Vectors are one-dimensional arrays that can hold numbers, words, or logical data (Kabacoff, 2011; p. 24). Note that vector elements can have only one mode i.e. be data of one type only.
You can create a vector using the concatenate or combine function c().
# a vector of numbers
A <- c(1,2,3,4,5)
# a vector of character strings
B <- c("a", "b", "c", "d", "e")
# a vector of logical values
C <- c(TRUE, TRUE, TRUE, FALSE, FALSE)
Notice:
— I am putting the characters in quotation marks when I create vector B. You will see everywhere in R that character strings (from elements of data, as in the example, to plot titles) are entered using either matching double (“) or single (‘) quotes but are printed using double quotes or sometimes without quotes (see here).
You can create vectors using also the functions seq() and rep(). You use seq() to get a sequence of numbers separated by equal steps
D <- seq(1,5) E <- seq(100,500,100)
Notice:
— The first function call, C <- seq(1,5), asked for a sequence of numbers from 1 to 5 in steps of 1: the default step size of 1 is used unless we add an argument to the seq() function call.
— the second call, D <- seq(100,500,100), asked for a sequence of numbers from 100 to 500 in steps of 100.
We could alternatively do the first kind of operation using the : operator e.g. e <- 10:50.
Notice:
— The seq() function affords greater flexbility.
The replicate rep() function allows you to create a generate repeated values. You might want to use this function if you were generating some coding values for a dataset. Let’s say you had a set of five observations all of which had been recorded by a researcher named RA, you could add a variable coding for the researcher’s identity by using rep(). What if you wanted to code for the presence of a feature, the feature is either there (code as 1) or not (code as 0) and it is there for the first three but not the second observations, you can create a vector of numbers doing the coding also using rep().
RA.ID <- rep("RA", 5)
coding <- rep(1:0, c(3,2))
Vectors are objects and you can interact with them. You can refer to specific elements in a vector using a vector.name[] position notation.
I mean, you can ask what the fourth element in a vector is by asking for it with C[4], or you can ask what the first and second elements in a vector are by asking for them with B[c(1,2)].
C[4] B[c(1,2)]
Notice:
— You will see c(), seq(), [], over and over again in R code.
Matrices
A matrix is a two-dimensional array where each element has the same mode, whether that be number, character or logical. I will be brief here because matrices are important to the statistical analyses you will use, and thus in how R works, but will not often be created directly by you.
Matrices can be created with the matrix(), cbind() and rbind() functions. I use the latter all the time so you’ll see them again. We might use matrix() again, I suspect for plotting model predictions, so need to at least lay out the basics here.
When using matrix(), you define a vector of values, then define how you want the matrix structured – how many rows, how many columns – then you define how you want the values put into the matrix structure; the default is by columns, but you could have it by rows.
# create a vector of numbers F <- seq(1,20) # create the matrix, with 5 rows and 4 columns G <- matrix(F, nrow = 5, ncol = 4) # same numbers, but different dimensions H <- matrix(F, nrow = 4, ncol = 5) # same numbers, 5 rows and 4 columns, but filled by rows rather # than by (default) columns I <- matrix(F, nrow = 5, ncol = 4, byrow = TRUE)
Try these bits of code out.
You can also create matrices by sticking vectors together using the column bind cbind() and row bind rbind() functions.
As noted, you will use these functions over and over again because you can bind not only vectors – as in these toy examples – but big, complicated, datasets (dataframes) using the exact same functions. It beats copy and paste in excel or SPSS.
We can exemplify the use of the functions with the vectors we have already created.
You can rbind() the vectors D and E together to create a matrix J with 2 rows and 5 columns. You can cbind() the vectors D and E together to create a matrix K with 5 rows and 2 columns
J <- rbind(D,E) K <- cbind(D,E)
Try these bits of code out.
Obviously, here, R does not know whether the vectors are ‘horizontal’ or ‘vertical’: both are just a string of numbers which could be either so the rbind() and cbind() calls are what determine whether you get a 2 x 5 or a 5 x 2 matrix of values.
Matrices have dimensions i.e. in the examples G is 5 x 4 matrix while H is 4 x 5.
The rbind() and cbind() functions will not succeed if the numbers of elements do not match, whether by row or by column, and you will get an error message tell you that the number of columns or rows must match, as you’ll see if you try this incorrect function call:
L <- rbind(G,H)
That will get you the error: ” Error in rbind(G, H) : number of columns of matrices must match (see arg 2) ”
Thus, to get rbind() to work: make sure that the number of columns for the matrices you are trying to bind do match, or make sure that the vectors you are trying to bind have the same number of elements, or that the vector you are trying to bind to a matrix has the same number of elements as the matrix has columns.
Obviously, the same restriction will apply to using cbind().
I can rbind() correctly the G and I matrices because they both have the same number of columns since they are two 5 x 4 matrices. I can cbind() the G matrix and the E vector because E has as many elements as G has rows.
L <- rbind(G,I) M <- cbind(G, E)
Notice:
— We cannot add values of a different data mode i.e words to a number matrix, as in:
N <- cbind(M, RA.ID)
— The function call will work but everything in the matrix will now be treated as a word or character string.
— If you want to have numbers and words in the same data table you will be working with dataframes, which we will focus on in the next post, and treat briefly here.
Matrices and referring to specific elements using subscripts
Here comes something really useful, that will remain useful throughout your R career.
You can identify rows, columns, or elements of a matrix by using the [] subscript notation available in R (Kabacoff, 2011; p. 25).
I cannot over-emphasize how handy this is.
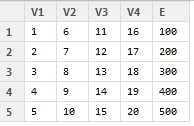
We can take one of the matrices we have built as an example in the foregoing, M. In creating M we put together a 5 x 4 matrix of numbers from 1:20 (G) with a vector of numbers from 100 – 500 (G). It looks like this:
Notice:
— I did not bother to name the rows or columns of the 5 x 4 matrix G, so R has supplied column names V1-4 and row names 1-5; obviously, vector E will get its name carried with it into the matrix M I created through cbind().
Anyway, for matrix M, I can refer to specific elements of the matrix by position using a subscript notation.
X[i,] refers to the ith row of matrix X.
X[,i] refers to the jth column of matrix X.
X[i,j] refers to the element at the ith row and jth column of X.
You could refer to multiple elements by entering a vector.
X[i, c(1,3)] refers to the 1st and 3rd elements at the ith row of X.
Now see how we can find out to use this notation to select – have reported to us – specific elements in a matrix.
# get the 2nd row in M M[2,] # get the second column in M M[,2] # get the element in the second row, second column in M M[2,2] # get the elements 1 and 3 in the second row in M M[2,c(1,3)]
Run the code and verify that the answers are correct.
Dataframes
I have mentioned dataframes before, here and elsewhere. As I have noted, they are what we will be dealing with most of the time, and they will appear most familiar to you, similar to the tables or sets of data you have used in SPSS or excel. You will know, from your experience, that you can get all different kinds of words into excel and (with restrictions i.e. specifying variable data types) into SPSS.
A dataframe is like a matrix. There are rows and columns of values. You can specify or work with specific elements in the dataframe using the [i rows, j columns] notation. You can work with dataframes using the cbind() and rbind() functions.
A dataframe is more general than a matrix because different columns can hold different kinds of data i.e. include variables with number values, or character etc. values.
You can create a data.frame using the data.frame() function.
Thus, if we just cbind() a matrix M of numbers and a vector RA.ID of character strings, we produce a matrix of character strings (all numbers get treated as characters by R).
While if we use data.frame() to put M together with RA.ID, we get a set of observations including both a vector of characters and a set of numbers.
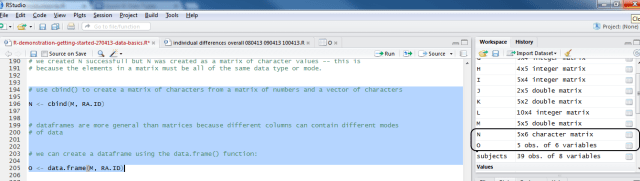
# use cbind() to create a matrix of characters from a matrix # of numbers and a vector of characters N <- cbind(M, RA.ID) # we can create a dataframe using the data.frame() function: O <- data.frame(M, RA.ID)
And you can see the difference in the way that R treats the objects N and O in how they are listed in the workspace window in R-studio, N as a matrix of characters, and O as a data.frame of 5 observations with 6 variables.
Notice: the difference in the listing on the right here.
What have we learnt?
[The .R script used in the examples can be downloaded from here.]
We have learnt about data structures, especially vectors, matrices and dataframes.
We have learnt how to create vectors, and how to refer to specific elements in a vector by position.
We have learnt how to create matrices, and how to refer to specific elements by position using the [] notation.
We have learnt about dataframes as a more general form of matrix.
Key vocabulary
c()
seq()
: operator
rep()
cbind()
rbind()
matrix()
[,] notation
data.frame()
Reading
The things treated here are very useful, so bear further study. There are clear chapters on this material in Kabacoff (2011) and Dalgaard (2008) – books that I recommend – among other sources.